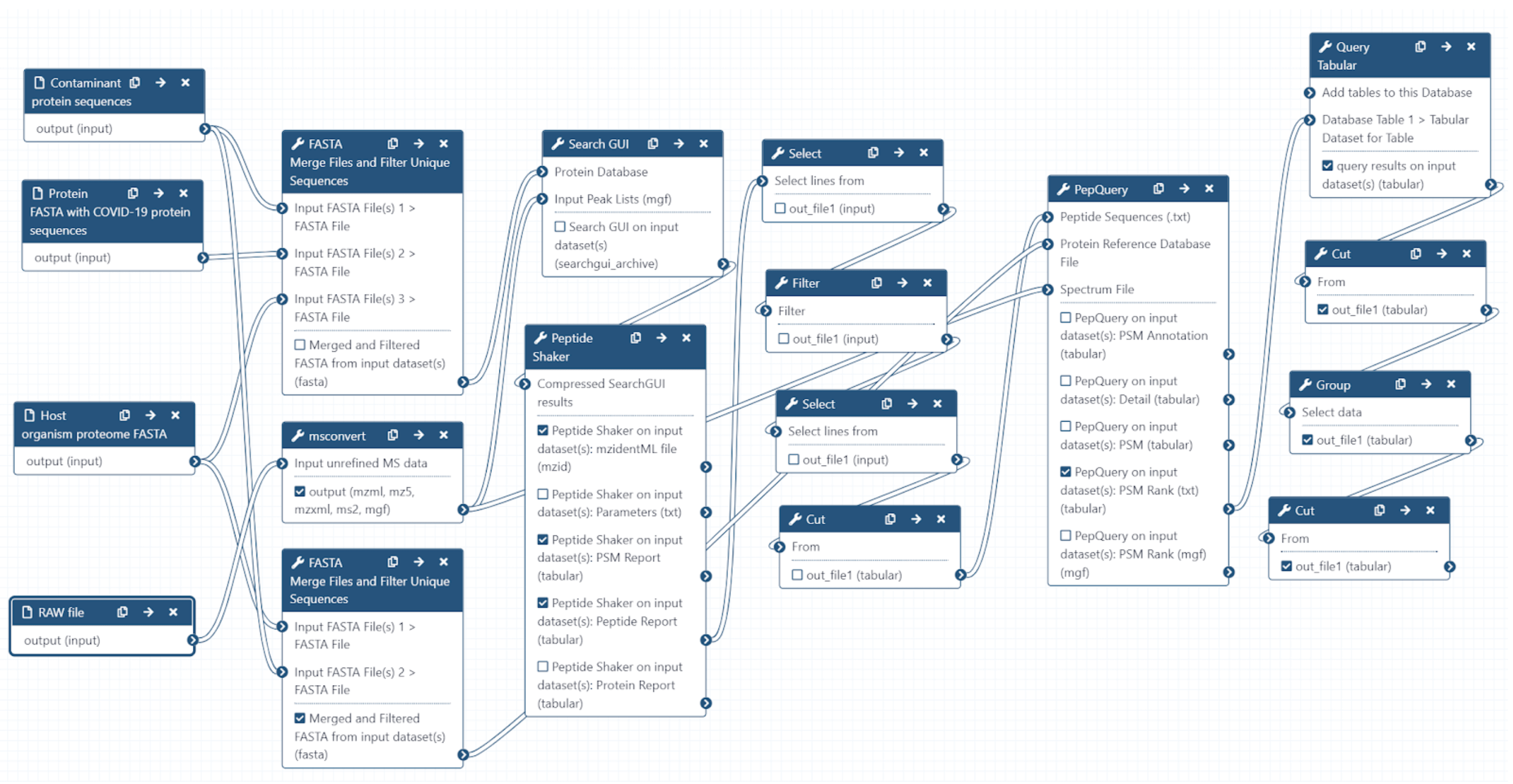
SEARCHGUI PROTEOMICS SOFTWARE
Although commercial and actively-supported software offers reliability and ease of use, its usage comes with a cost and usually includes canned features that are used for most standard datasets. LFQ analysis can be performed by public domain software suites such as MaxQuant and Skyline, or by commercial software such as PEAKS and Progenesis. Currently, there are several software packages available for LFQ analysis. LFQ is a useful method for quantification when the introduction of stable isotopes is impractical (for example, in human or animal model studies) or for applications such as proteogenomics or metaproteomics, which rely on peptide-level quantification. In the case of the label-free quantification (LFQ) methods, the peak intensity or area under the curve of a detected peptide ion allows the relative quantification of peptides across different samples. This work establishes a process for software implementation, optimization, and validation, and offers access to two robust software tools for LFQ-based analysis within the Galaxy platform. Software features evaluated include: (a) match-between-runs (MBR) (b) using multiple file-formats as input for improved quantification (c) use of containers and/or conda packages (d) parameters needed for analyzing large datasets and (e) optimization and validation of software performance. Through rigorous testing and communication with the tool developers, we have optimized the performance of each tool.
In this study, we have evaluated moFF and FlashLFQ, two open-source LFQ tools, and implemented them within the Galaxy platform to offer access and use via established workflows. However, workflows within the Galaxy platform have lacked well-tested LFQ tools.

Bioinformatics workflows accessible via the Galaxy platform have proven useful for analysis of such complex multi-omic studies. LFQ enables peptide-level quantitation, which is useful in proteomics (analyzing peptides carrying post-translational modifications) and multi-omics studies such as metaproteomics (analyzing taxon-specific microbial peptides) and proteogenomics (analyzing non-canonical sequences). For mass spectrometry-based peptide and protein quantification, label-free quantification (LFQ) based on precursor mass peak (MS1) intensities is considered reliable due to its dynamic range, reproducibility, and accuracy.
